- BioPharm International-08-02-2010
- Volume 2010 Supplement
- Issue 6
Protein Characterization Through the Stages
Biomanufacturers should follow a risk-based approach to decide which methods to use to characterize their products.
ABSTRACT
Protein characterization plays a critical role in determining the safety and efficacy of biological products. Advances in analytical techniques have given biopharmaceutical companies new methods for characterizing their products; the challenge has become choosing the right combination of methods. Biomanufacturers must leverage their experience to determine which methods to apply at each stage of the drug development process, and use a risk-based approach to guide their selection.
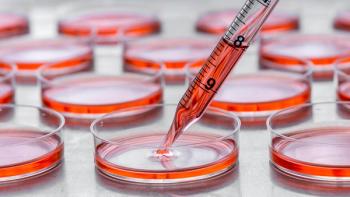
Photo courtesy of: DASGIP
Protein characterization plays a critical role in the discovery, development, and manufacture of biopharmaceutical products. Unlike traditional pharmaceutical companies, biopharmaceutical manufacturers must rely on complex biological systems to produce their therapeutic products. The inherent variability associated with biological expression systems necessitates comprehensive characterization of the final product to ensure comparability throughout the development process and from batch to batch post-approval.1 The advent of biosimilars has given protein characterization further relevance as it forms the foundation for assessing relative similarity. Advances in analytical techniques have given biopharmaceutical companies new and increasingly sensitive methods for characterizing their products; the challenge has become choosing the right combination of methods, and balancing the risk of not detecting a material change in the product with the time and expense associated with extensive characterization.
Role of Protein Characterization at Various Stages of Bioprocessing
Protein characterization data play different roles at various stages of drug discovery, development, and manufacturing. At the discovery stage, these data form the basis for candidate screening and can provide valuable guidance on cell line and strain selection and upstream operating conditions. Assays used at this stage should be focused on determining relative yields from various cell lines or strains, examining post-translational modifications, identifying lead candidates with desirable properties (e.g., high in vitro enzyme activity, solubility at physiological pH), and screening out molecules that raise red flags regarding potential immunogenicity and half-life issues (e.g., readily oxidized or deamidated molecules).2 These analyses also can provide a preliminary assessment of short-term stability at the target pH, salt concentration, and storage conditions.
As molecules progress through the development process, protein characterization techniques are used to guide process optimization—identifying operating conditions that produce higher yields and increased purity, and limit aggregation. The data obtained also are used to support formulation development, to develop product storage and handling instructions, and to define product specifications.3 Relevant analytical methods are used for in-process monitoring of intermediates, final product analysis, and reference standard characterization in preparation for preclinical and clinical manufacturing. As biopharmaceutical products move into clinical manufacturing, detailed analyses allow systematic collection of stability data, demonstrate comparability following scale up or a manufacturing site change, and during process validation. Novel assays and increasingly sensitive analytical methods can identify minor changes in protein structure, composition, or impurity profiles which could have significant effects on in vivo immunogenicity, toxicity, and half life.4 The FDA recommends that manufacturers use orthogonal methods for analyzing biopharmaceutical product purity and identity, so a combination of different techniques must be selected that provide a balance between regulatory compliance and commercialization.5
This balance can be difficult to define, however. The FDA has advocated the use of a risk-based approach for evaluating product comparability subsequent to a manufacturing change (e.g., scale up, site transfer, process changes). This involves the manufacturer engaging in a theoretical analysis of how the change could potentially affect both the impurity profile and the product itself, and then designing a characterization program which enables detection of these "high probability" changes as well as any "high impact" changes that would significantly affect product quality, safety, or efficacy should they occur.6 Companies can leverage their experience with biological products in general and with the specific molecule in particular to conduct these comprehensive risk assessments. A change identified as high risk, i.e., likely to result in material changes to the final product, would then lead to extensive testing to determine comparability between the product pre- and post-change. Examples of high-risk changes would include moving from serum-containing to serum-free media upstream or moving from crystallization to chromatography downstream. Low-risk changes such as using a new supplier of a common buffer would dictate a less comprehensive evaluation.
Current State of the Science
Existing characterization methods are intended to provide a detailed, comprehensive analysis of protein size, charge, purity, activity, and structure (primary, secondary, and tertiary). These analyses also are used to examine impurity profiles, focusing particularly on aggregates, a key concern for regulatory agencies because of their potential immunogenicity.7 A broad range of analytical methods can be used to characterize proteins on the basis of identity, purity, yield, aggregation, specificity, and activity. These methods are used to assess comparability, form the basis for product release, determine product stability, and guide formulation development. Table 1 summarizes the most commonly used characterization methods for biopharmaceutical products.
Table 1. Most commonly used characterization methods for biopharmaceutical products
Selecting the specific analytical methods to be used in characterizing a given biomolecule involves an assessment of the capabilities, advantages, and limitations of the available options. The sensitivity of the method in question is critical to identify minor contaminants or product variants that could potentially elicit significant immune responses in vivo. Reverse phase high performance liquid chromatography (RP-HPLC), mass spectrometry (MS), and dynamic light scattering (DLS) provide high sensitivity and are commonly used throughout the drug discovery-development-manufacturing chain. Size exclusion chromatography (SEC), analytical ultracentrifugation, and UV-Vis spectrophotometry are valued for their ability to accurately quantify specific analytes. Specificity is best determined with biological assays that mimic the desired in vivo molecular interactions in vitro, such as Western blots, enzyme assays, and cell-based assays.8 From both an operational and quality perspective, the best assays are robust, relatively insensitive to changes in sample matrix or product concentration, rapid, and easy to perform.
Recent Trends
No single assay possesses all of the attributes described above, but improvements in existing methods and the development of novel technologies have provided the biopharmaceutical industry with better tools for protein characterization. With the advent of biosimilars, regulatory agencies have become more rigorous in their analyses of comparability in terms of protein structure and have demonstrated notable interest in higher order structures and protein aggregates. As a result, biomanufacturers are incorporating more comprehensive structural analyses into their product characterization assay portfolios.
Higher Order Structures
A number of techniques have emerged as valuable tools for evaluating higher order structures of protein biopharmaceuticals, including x-ray crystallography, light scattering, calorimetry, and spectroscopy. Advanced spectroscopic techniques such as nuclear magnetic resonance (NMR), circular dichroism (CD), and fluorescence spectroscopy (FS) have proven particularly useful for structural analysis.
CD is a form of UV absorption spectroscopy, and typically is used to examine the secondary structure of proteins. CD often is used to assess the conformational stability of a protein under stress, providing insight into a protein's stability at various temperatures and pH levels and in the presence of denaturing agents. Using this method, appropriate buffer compositions, stabilizers, and excipients can be identified to increase the melting temperature or the reversibility of thermal unfolding for a given protein, which in turn result in enhanced shelf life for the final drug product. Circular dichroism also can determine whether protein–protein or protein–ligand interactions have the potential to alter the conformation of the target protein, as conformational changes produce a spectrum which differs from the sum of the individual components. CD is a very sensitive technique for analyzing a protein's secondary structure, requiring only µLs of solution at concentrations as low as 50 µg/mL protein, and the sample can be analyzed in any buffer that does not have a high absorbance in the far-UV region of the spectrum.9
Fluorescence spectroscopy, a form of UV excitation spectroscopy, is commonly used to study the tertiary structure of proteins. The intrinsic fluorescence of a given molecule is determined by the presence and location of tryptophan, tyrosine, and phenylalanine residues in the target molecule.10 Proteins also can be labeled with fluorophores and the extrinsic fluorescence used to determine tertiary structure. A commonly used covalent probe is fluorescein 5'-isothiocyanate, which can be attached to lysine residues in the target protein. Fluorescence spectroscopy is accurate, highly sensitive, and can be used even at very low concentrations of the target protein.
Another technique that has recently gained attention for its potential applications in higher order structure analysis is hydrogen-deuterium exchange with mass spectrometry (H/DX–MS). This method exploits the natural tendency of hydrogen atoms in a protein to exchange with hydrogen atoms in the surrounding solvent.11 If an isotope of hydrogen is used as the solvent, namely deuterium oxide, its heavier mass gets incorporated into the protein. Because the protein now weighs more than normal, this change in mass can be monitored with high-resolution mass spectrometers. The rate of exchange of hydrogen atoms is used to determine the protein structure. Compared with other techniques for analyzing higher order protein structures, H/DX–MS is extremely sensitive, requiring only picomole quantities of protein. Further, the technique can be applied to proteins of varying sizes, including large protein complexes. Perhaps most notably, the technique can effectively analyze membrane proteins, which are very difficult to examine with most other protein characterization techniques.
Aggregates
Aggregates have been identified as having a potential causal link to increased immunogenicity in vivo. As a result, biomanufacturers are under increasing pressure from regulatory bodies to provide detailed information about the quantity and nature of any aggregates present in a biopharmaceutical product. Aggregation can be particularly challenging to evaluate, as it can occur during manufacture, storage, and handling. Information on aggregate formation therefore can be of particular value during the development process in evaluating possible formulations for the drug product. Aggregate concentration also is a critical parameter to be monitored in stability studies.
Sedimentation velocity analytical ultracentrifugation (AUC–SV) uses strong centrifugal force to separate various species in a given sample mixture. This method can be used to study samples over a relatively wide range of pH and ionic strength conditions, and at temperatures from 4 to 40 °C.12 Sample volumes are typically <1 mL, and the total mass of protein required is <1 mg. AUC–SV typically is used as an orthogonal method to SEC, because of its accuracy, high resolution of aggregates, and ability to analyze aggregates in a wide range of buffers. The technique is time-consuming, however, and requires specialized equipment, so it has not yet gained acceptance as a standard test method for product release.
Asymmetric field flow fractionation (aFFF) also uses non-column technology to characterize aggregates, and therefore is also appropriate for use as an orthogonal method to SEC. This method uses a semi-permeable membrane and two perpendicular fluid flows in a channel to separate macromolecules based on molecular weight and hydrodynamic size.13 The technique is gentler to macromolecules than SEC, and is therefore less likely to change aggregate composition during the analysis. Also, aFFF has a wider dynamic range than SEC, enabling detection of larger aggregates. The method does have limitations, including the potential for molecular interactions between the proteins and the membrane, as well as a lack of precision. Used in conjunction with other complimentary methods, aFFF can form part of a comprehensive evaluation of protein aggregation.
Summary
As the biopharmaceutical industry continues to evolve, so must its analytical techniques. Protein characterization plays a critical role in determining the safety and efficacy of biological products, and as such there is a need to continue to improve the accuracy, sensitivity, specificity, and robustness of protein assays, and to develop novel assays that target specific areas of concern. Biomanufacturers must leverage their experience and knowledge to determine which methods to apply at each stage of the drug development process, and use a risk-based approach to guide their selection. Collaboration between manufacturers and regulatory agencies is critical in the ongoing search for an answer to the question—How well characterized is well characterized enough?
Lisa Crossley, PhD, the founder and principal of BioVentures, Ontario, Canada, 888.405.9549,
References
1. Morrow KJ. Tools for protein structure characterization. Gen Eng Biotech News. May 2008;28(9).
2. Magil S. Biopharmaceutical characterization techniques for early phase development of proteins. BioPharm Int. Guide to Bioanalytical Advances. Sept 2005; p. 34–42.
3. Towns J, Webber K. Demonstrating comparability for well-characterized biotechnology products: Early phase, late phase, and post-approval. BioProcess Int. Feb 2008:6(2):32–43.
4. Krishnamurthy R, Sukumar M, Das T, Lacher N. Emerging analytical technologies for biotherapeutics development. BioProcess Int. May 2008;6(5):32–42.
5. US Food and Drug Adminsitration. Guidance for Industry. Q6B specifications: Test procedures and acceptance criteria for biotechnological/biological products. Rockville, MD; Aug 1999.
6. Dougherty J, et al. Postapproval changes for large-scale biopharmaceutical manufacturing: Global regulatory issues. Langer ES, ed. In: Advances in large-scale biopharmaceutical manufacturing. ASM Press: Washington, DC, 2004; pp. 555–91.
7. Patten PA, Schellekens H. The immunogenicity of biopharmaceuticals: lessons learned and consequences for protein drug development. Brown F, Mire-Sluis AR, editors. Immunogen Therapeutic Bio Prod.
8. Aldridge S. Next-generation protein characterization. Gen Eng Biotech News. Apr 2006;26(7).
9. Lipp, E. Characterizing Proteins to Bolster Pipelines. Genetic Eng Biotech News. June 2010; 30(12).
10. Vivian JT, Callis PR. Mechanisms of tryptophan fluorescence shifts in proteins. Biophys J. 2001;80(5):2093–109.
11. Wales TE, Engen JR. Hydrogen exchange mass spectrometry for the analysis of protein dynamics. Mass Spectrom Rev. 2006;25(1):158–70.
12. Berkowitz SA. Role of analytical ultracentrifugation in assessing the aggregation of protein biopharmaceuticals. AAPS J. 2006;8:590–605.
13. Gabrielson JP, et al. Quantitation of aggregate levels in a recombinant humanized monoclonal antibody formulation by size-exclusion chromatography. Asymmetrical flow field flow fractionation, and sedimentation velocity. J Pharm Sci. 2007;96:268–79.
Articles in this issue
over 15 years ago
BioPharm International, August 2010 Supplement (PDF)over 15 years ago
Biophysical Characterization for Product ComparabilityNewsletter
Stay at the forefront of biopharmaceutical innovation—subscribe to BioPharm International for expert insights on drug development, manufacturing, compliance, and more.