Manufacturing, Cell Therapies
Latest News
Latest Videos

More News
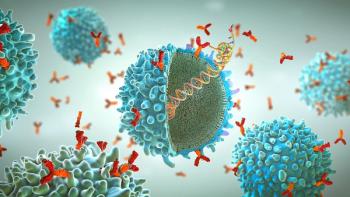
Single-use technologies provide efficiency and flexibility while reducing risk in the manufacture of cell and gene therapies.
The company hopes the rebranding effort will better align with its work in cell culture media manufacturing.

Key 2025 FDA draft and final guidances emphasize the modernization of biotech regulations, acceleration of rare-disease therapies, and streamlining of biosimilar pathways.

Charles River has launched a second cohort to speed CGT innovation with technical guidance and scalable manufacturing support.

Flexible manufacturing processes and facilities support the pipeline of allogeneic cell therapies.

How is the biopharma industry solving manufacturing and scale-up bottlenecks for cell and gene therapies? Read on to find out.
A new Cue Biopharma–ImmunoScape partnership seeks to advance targeted TCR-T expansion for solid tumors, supporting broader access and improved clinical durability.

The partnership is aiming to embed digital identification into labware, solving chain-of-identity challenges for individualized cell and gene therapies.

As cell and gene therapy is poised to play a major role in the future of medicine, it’s important to know the basics.
Jon Ellis, CEO, shares his thoughts on Trenchant BioSystems’ new technology, its reception within the cell and gene therapy sector, and the future nature of industry partnerships.

In a session at the Cell and Gene Meeting on the Mesa, Prime Medicine CEO Allan Reine discussed how prime editing offers versatile, safe gene correction, but that delivery to target cells remains a major hurdle.

A panel at the Cell and Gene Meeting on the Mesa discussed how advanced therapy production demands modular platforms, automation, and data governance to drastically improve patient access and affordability.

The new protein-based HDR enhancer aims to improve CRISPR precision for advancing cell and gene therapy development workflows.

Engineered biosynthetic pathways provide new avenues for accessing complex molecules.

A collaboration between Limula and Institut Paoli-Calmettes aims to advance automated stem cell transplant processing to improve cryoprotectant removal, enhance patient outcomes, and streamline manufacturing.
Allogeneic CAR-T therapies deliver scalable, off-the-shelf cancer therapy, while autologous CAR-T therapies provide patient-specific but time-intensive treatment.

ElevateBio BaseCamp Achieves First Multi-Modality ICMC Certification in Commercial CGT Manufacturing
Third-party ICMC certification verifies multi-modality manufacturing readiness, meeting US and EU standards for advanced genetic therapies.
The Hopewell, N.J., site adds scalable, end-to-end viral vector production with integrated quality systems to speed clinical and commercial gene therapy programs.

The newly launched facility is located in The Woodlands, Texas, and will produce plasmid DNA as well as strengthen biopharma supply chains.

Minimize contamination in cell therapy manufacturing with isolators, staff protocols, sterile materials, and validated suppliers for process integrity.

The consortium will focus on the delivery of a fully automated robotics cell and gene therapy manufacturing platform.

BioPharm International® spoke with Jon Ellis, CEO, Trenchant BioSystems, about the commercial landscape for cell therapies and how digitization is impacting the industry.

The industry is diversifying pipelines from traditional small-molecule drugs to embrace complex and exciting new modalities.

Global scale-up requires developing replicable processes that work the same no matter where they are performed. This can be accomplished with smart factories that utilize fully automated manufacturing platforms.

The company’s technology was used to create CAR-T cells that demonstrated the expression of complex CARs in a single DNA donor.